boiler noise
Dept. of Medical Biostatistics,
Dept. of Mathematics & Statistics,
College of Engineering & Mathematical Sciences,
University of Vermont
Office: Innovation Hall, E437
Publications: Google Scholar
Curriculum Vitae
Research Focus
![]()
Statistical Genetics Projects:
-
Software projects
IDAWG The Immunogenomics Data-Analysis Working Group
Links to some collaborative work.
Director of the Health Outcomes Research Internship Program (HORIP) with the Vermont Department of Health (VDH)
Statistics and Biostatistics Journal Club
![]()
My research in the area of statistical genetics has been in human population, disease, and evolutionary genetics, with an emphasis on the genes in the human leukocyte antigen (HLA) region. The genes in this region are highly polymorphic (over 400 alleles identified for some genes) and control several functions in the immune response. HLA genes influence susceptibility to over 200 diseases. These diseases include autoimmune diseases (e.g., type 1 diabetes, multiple sclerosis, and rheumatoid arthritis), cancers (e.g., Hodgkin disease), and infectious diseases (e.g., tuberculosis, malaria, and AIDS). They are genetically complex in that they involve a number of genes and genetic heterogeneity. Several functional and population studies have indicated that the extensive variation at HLA loci is due to natural selection.
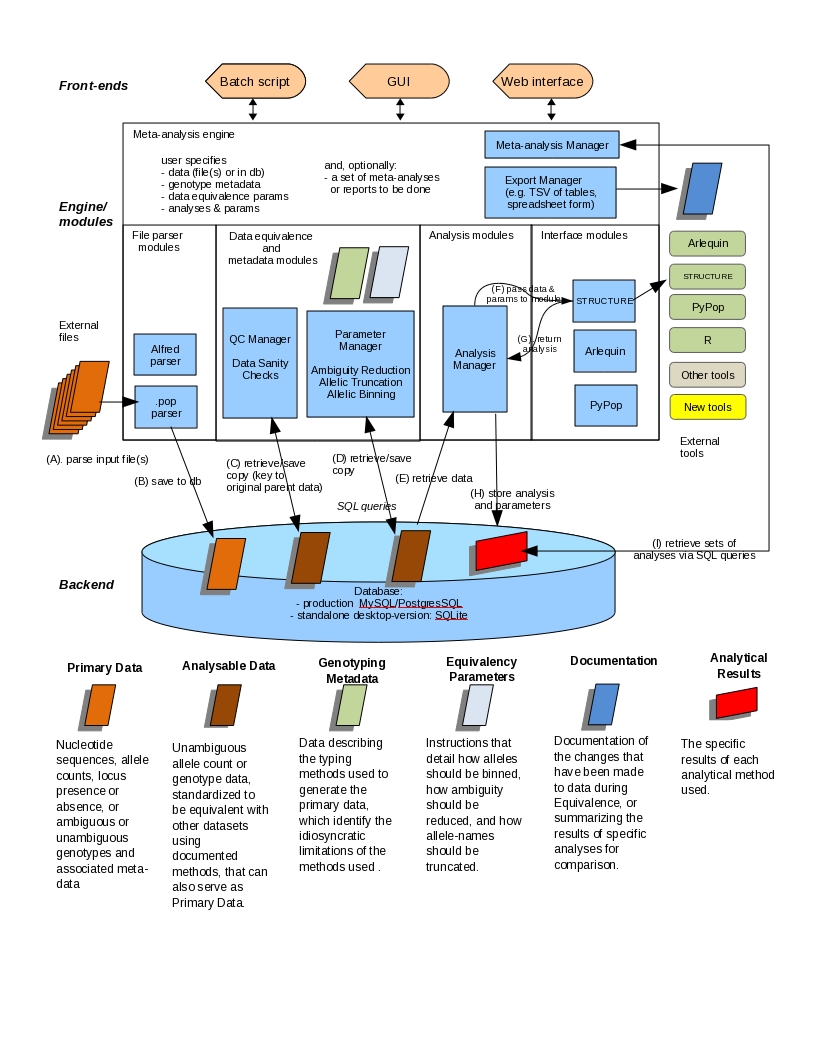
My main research goals are to elucidate the evolutionary forces acting on the HLA region, to distinguish between selective and demographic pressures, and to determine the role of HLA genes in disease and human stem cell transplantation outcomes, including the identification of additional genetic factors involved. Specific areas of research include determining the effect of errors in haplotype frequency estimation on the type I error rate and on power in case-control studies, investigating the properties of permutation tests for linkage disequilibrium at the individual haplotoype level, and determining the properties of several measures of linkage disequilibrium. I have been working on a software development project, PyPop: A Software framework for Population Genomics, which recently had a second beta release. This software program has been a necessary tool to address the aforementioned research goals.
![]()
Bios of some famous Statisticians
![]()